library(tidyverse)
library(broom)
library(quantreg)
library(marginaleffects)
library(here)Chapter 6: Regression with One Binary and One Numeric Predictor
1 Introduction
In the previous chapter, we studied regression models with two binary predictors, and learned how to distinguish between:
- additive models (no interaction)
- interaction models (effect modification)
In this chapter, we extend these ideas to a model with:
- one binary predictor:
trt(therapist-guided vs waitlist) - one numeric predictor:
phq9_screen(baseline depression severity)
This allows us to introduce conditional effects when the moderator is numeric, and to distinguish between:
- parallel slopes (no interaction): the treatment effect is constant across
phq9_screen - effect modification (interaction): the treatment effect varies with
phq9_screen
You will learn:
- how to interpret models with one binary and one numeric predictor
- how conditional treatment effects vary across values of a numeric variable
- how interactions change interpretation
- how to estimate conditional and marginal effects using
marginaleffects - how to visualize interactions with predicted values
2 Packages and Data
2.1 Load data
df_clean <- read_csv2(here("data", "steps_clean.csv"))As before, we use treatment group as our binary predictor. We subset the data to include only participants in the therapist-guided and waitlist conditions.
df_red <- filter(df_clean, trt %in% c("therapist-guided", "waitlist"))
df_red <- df_red |> mutate(
trt = factor(trt, levels = c("waitlist", "therapist-guided"))
)The other predictor will be baseline depression severity measured continuously using phq9_screen.
summary(df_red$phq9_screen) Min. 1st Qu. Median Mean 3rd Qu. Max.
1.00 7.00 9.00 9.85 13.00 19.00 3 One Binary and One Numeric Predictor without Interaction (Additive Model)
We begin with a model that includes both predictors without an interaction.
3.1 Model formula
lsas_post ~ trt + phq9_screenThis corresponds to the linear regression model for the conditional mean:
\[ \begin{aligned} \mathbb{E}(LSAS_{post,i} \,\mid\, trt_i,\; phq9\_screen_i) =& \beta_0 + \\ & \beta_1\, \mathbf{1}\{\text{trt}_i = \text{therapist-guided}\} + \\ & \beta_2\, phq9\_screen_i \end{aligned} \]
Where:
- \(\beta_0\): mean outcome when
trt = waitlistandphq9_screen = 0 - \(\beta_1\): conditional treatment effect holding PHQ-9 constant
- \(\beta_2\): conditional effect of PHQ-9 holding treatment constant
In this model:
- the treatment effect is assumed to be the same at all values of PHQ-9
- the PHQ-9 slope is assumed to be the same in both treatment groups
This is an additive model (parallel slopes).
4 Linear Regression without Interaction
4.1 Fit the model
mod_lm_add <- lm(lsas_post ~ trt + phq9_screen, data = df_red)
summary(mod_lm_add)
Call:
lm(formula = lsas_post ~ trt + phq9_screen, data = df_red)
Residuals:
Min 1Q Median 3Q Max
-65.918 -13.561 -0.148 12.947 62.865
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 63.3772 5.9399 10.670 < 2e-16 ***
trttherapist-guided -19.3976 4.4906 -4.320 3.46e-05 ***
phq9_screen 1.4617 0.4919 2.972 0.00364 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 23.55 on 109 degrees of freedom
(8 observations deleted due to missingness)
Multiple R-squared: 0.2229, Adjusted R-squared: 0.2086
F-statistic: 15.63 on 2 and 109 DF, p-value: 1.074e-06- Intercept: mean LSAS at post when
trt = waitlistandphq9_screen = 0
(note:phq9_screen = 0may be outside the range of the data, so the intercept is often not directly meaningful) - trt coefficient: treatment effect adjusted for PHQ-9
- phq9_screen coefficient: association between baseline depression and post-treatment LSAS, adjusted for treatment
4.2 Predicted means for each treatment group (averaged over PHQ-9)
avg_predictions(mod_lm_add, by = "trt")| trt | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| waitlist | 78.4 | 3.09 | 25.4 | <0.001 | 72.4 | 84.5 |
| therapist-guided | 57.4 | 3.21 | 17.9 | <0.001 | 51.1 | 63.6 |
4.3 Treatment effect as a conditional contrast
Because there is no interaction, the conditional treatment effect is the same for all PHQ-9 values.
avg_comparisons(mod_lm_add,
variables = "trt",
by = "phq9_screen"
)| phq9_screen | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| 1 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 2 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 3 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 4 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 5 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 6 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 7 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 8 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 9 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 10 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 11 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 12 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 13 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 14 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 15 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 16 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 17 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 18 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
| 19 | -19.4 | 4.49 | -4.32 | <0.001 | -28.2 | -10.6 |
5 Adding an Interaction
Now we allow the treatment effect to differ depending on baseline depression severity.
5.1 Model formula with interaction
lsas_post ~ trt * phq9_screenWhich expands to the conditional mean:
\[ \begin{aligned} \mathbb{E}(LSAS_{post,i} \,\mid\, trt_i,\; phq9\_screen_i) =& \beta_0 + \\ & \beta_1\, \mathbf{1}\{\text{trt}_i = \text{therapist-guided}\} + \\ & \beta_2\, phq9\_screen_i + \\ & \beta_3\, \mathbf{1}\{\text{trt}_i = \text{therapist-guided}\} \times phq9\_screen_i \end{aligned} \]
- \(\beta_3\) is the interaction term, it captures how the treatment effect changes as PHQ-9 increases
We now allow the treatment effect to differ depending on baseline depression, rather than remaining constant across PHQ-9 values.
6 Linear Regression with Interaction
6.1 Fit the interaction model
mod_lm_int <- lm(lsas_post ~ trt * phq9_screen, data = df_red)
summary(mod_lm_int)
Call:
lm(formula = lsas_post ~ trt * phq9_screen, data = df_red)
Residuals:
Min 1Q Median 3Q Max
-65.585 -13.229 -0.181 13.429 62.900
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 65.4083 7.5650 8.646 5.45e-14 ***
trttherapist-guided -23.5820 10.5995 -2.225 0.0282 *
phq9_screen 1.2647 0.6691 1.890 0.0614 .
trttherapist-guided:phq9_screen 0.4324 0.9913 0.436 0.6636
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 23.64 on 108 degrees of freedom
(8 observations deleted due to missingness)
Multiple R-squared: 0.2243, Adjusted R-squared: 0.2027
F-statistic: 10.41 on 3 and 108 DF, p-value: 4.513e-06- The treatment effect is conditional on PHQ-9
- Main effects are interpreted at the reference level of the other variable:
- the
trtcoefficient is the treatment effect whenphq9_screen = 0 - the
phq9_screencoefficient is the PHQ-9 slope in the waitlist group (reference level)
- the
- The interaction coefficient represents how much the treatment effect changes for a one-unit increase in PHQ-9. For example, a negative interaction means the treatment effect becomes more negative (larger improvement) at higher PHQ-9.
6.2 Conditional treatment effects at selected PHQ-9 values
A simple way to interpret the interaction is to estimate the treatment effect at several meaningful PHQ-9 values (e.g., low, medium, high).
phq_vals <- quantile(
df_red$phq9_screen,
probs = c(.25, .50, .75),
na.rm = TRUE
)
phq_vals25% 50% 75%
7 9 13 avg_comparisons(
mod_lm_int,
variables = "trt",
newdata = datagrid(
phq9_screen = phq_vals
),
by = "phq9_screen"
)| phq9_screen | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| 7 | -20.6 | 5.23 | -3.93 | < 0.001 | -30.8 | -10.30 |
| 9 | -19.7 | 4.56 | -4.32 | < 0.001 | -28.6 | -10.76 |
| 13 | -18.0 | 5.58 | -3.22 | 0.00129 | -28.9 | -7.02 |
These estimates answer: What is the treatment effect at low, medium, and high baseline depression?
6.3 Marginal treatment effect
avg_comparisons(
mod_lm_int,
variables = "trt"
)| Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|
| Type: response | |||||
| -19.4 | 4.51 | -4.3 | <0.001 | -28.2 | -10.5 |
This is the marginal (population-average) treatment effect, obtained by averaging the conditional treatment effects over the observed distribution of PHQ-9 values in the sample.
7 Quantile Regression with One Binary and One Numeric Predictor
Now let’s do the same thing with quantile regression (median regression).
7.1 Additive model (median differences)
mod_rq_add <- rq(
lsas_post ~ trt + phq9_screen,
tau = 0.5,
data = df_red
)
summary(mod_rq_add)
Call: rq(formula = lsas_post ~ trt + phq9_screen, tau = 0.5, data = df_red)
tau: [1] 0.5
Coefficients:
coefficients lower bd upper bd
(Intercept) 64.66667 60.36987 78.82968
trttherapist-guided -22.75000 -29.61235 -17.25753
phq9_screen 1.41667 0.04756 3.421087.2 Interaction model (median differences)
mod_rq_int <- rq(
lsas_post ~ trt * phq9_screen,
tau = 0.5,
data = df_red
)
summary(mod_rq_int)
Call: rq(formula = lsas_post ~ trt * phq9_screen, tau = 0.5, data = df_red)
tau: [1] 0.5
Coefficients:
coefficients lower bd upper bd
(Intercept) 62.00000 49.56531 79.43182
trttherapist-guided -20.08333 -40.19799 -9.85694
phq9_screen 1.75000 -0.28541 4.27356
trttherapist-guided:phq9_screen -0.33333 -1.78486 2.242217.3 Conditional median treatment effects at selected PHQ-9 values
avg_comparisons(
mod_rq_int,
variables = "trt",
newdata = datagrid(
phq9_screen = phq_vals
),
by = "phq9_screen"
)Warning: 2 non-positive fis| phq9_screen | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| 7 | -22.4 | 4.71 | -4.75 | < 0.001 | -31.7 | -13.18 |
| 9 | -23.1 | 4.71 | -4.90 | < 0.001 | -32.3 | -13.85 |
| 13 | -24.4 | 7.67 | -3.18 | 0.00146 | -39.4 | -9.38 |
7.4 Marginal median treatment effect
avg_comparisons(
mod_rq_int,
variables = "trt"
)Warning: 2 non-positive fis| Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|
| Type: response | |||||
| -23.3 | 5.03 | -4.63 | <0.001 | -33.2 | -13.5 |
8 Visualization
To visualize conditional effects when the moderator is numeric, it is useful to plot predicted values as a function of phq9_screen, separately by treatment group.
8.1 Predicted means across PHQ-9 (additive model)
plot_predictions(mod_lm_add, by = c("phq9_screen", "trt")) +
labs(
x = "PHQ-9 at screening",
y = "Estimated Mean LSAS (95% CI)",
color = "Treatment group",
fill = "Treatment group",
title = "Additive Model: Parallel Slopes (No Interaction)"
) +
theme_minimal()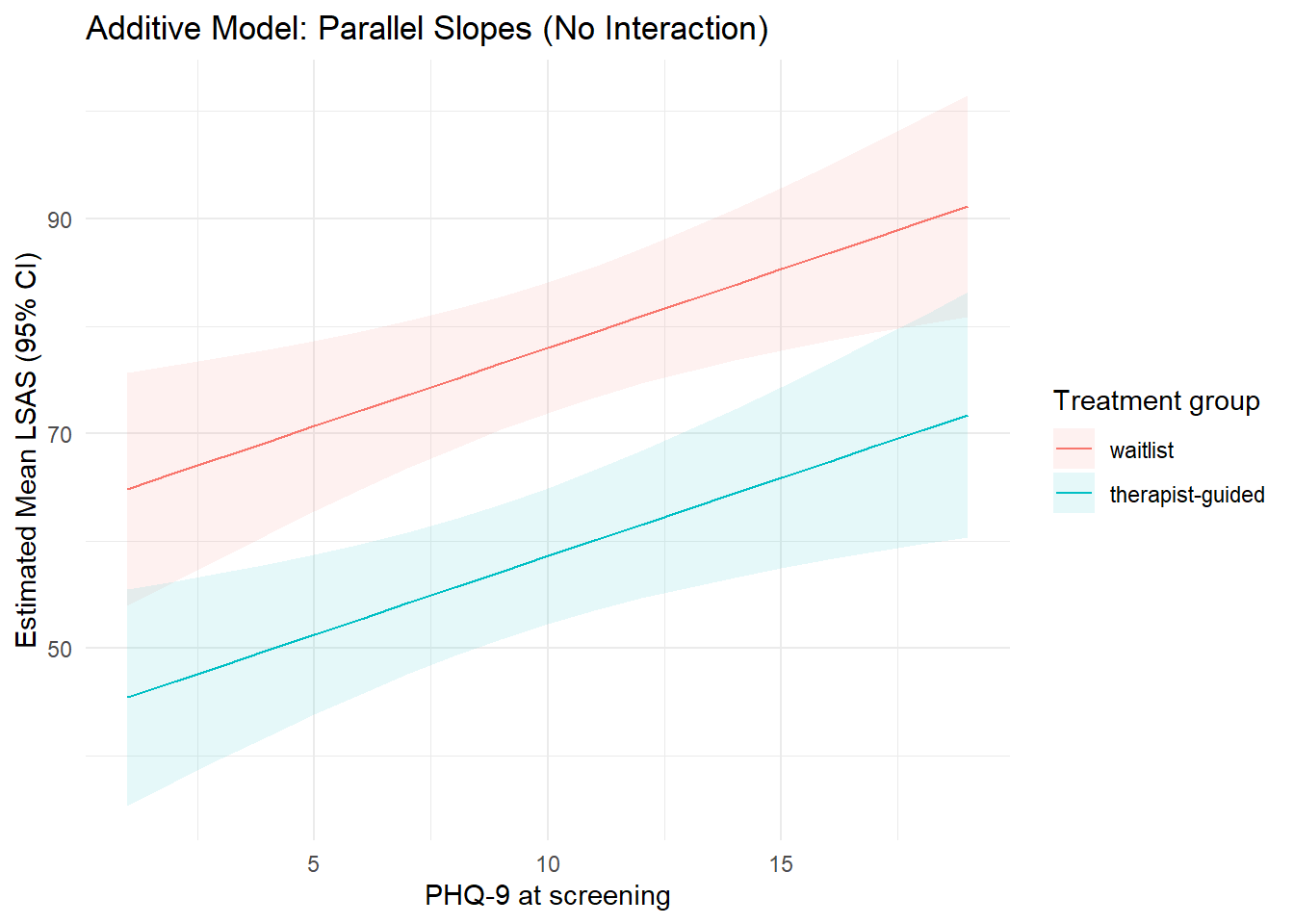
In the additive model, the lines are parallel: the difference between treatment groups is constant across PHQ-9.
8.2 Predicted means across PHQ-9 (interaction model)
plot_predictions(mod_lm_int, by = c("phq9_screen", "trt")) +
labs(
x = "PHQ-9 at screening",
y = "Estimated Mean LSAS (95% CI)",
color = "Treatment group",
fill = "Treatment group",
title = "Linear Interaction Model: Treatment Effect Varies with PHQ-9"
) +
theme_minimal()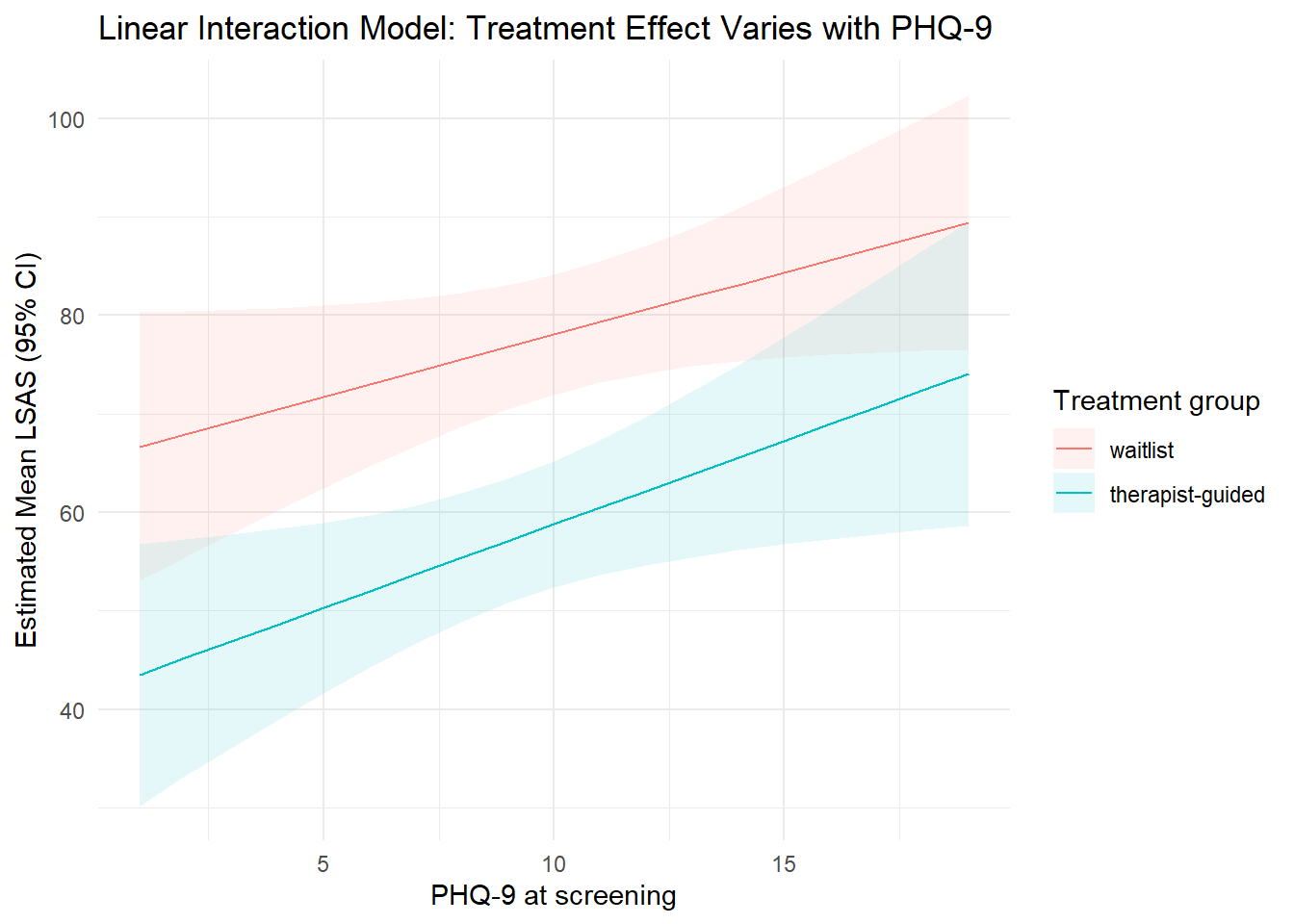
If the lines are not parallel, the treatment effect depends on baseline depression severity.
8.3 Visualizing the conditional treatment effect across PHQ-9
Instead of plotting predicted means, we can directly plot the estimated treatment effect as a function of PHQ-9, which makes it easier to gauge how much the treatment effect varies linearly with baseline PHQ-9 scores.
plot_comparisons(
mod_lm_int,
variables = "trt",
by = "phq9_screen"
) +
geom_hline(yintercept = 0, linetype = "dashed") +
labs(
x = "PHQ-9 at screening",
y = "Estimated Treatment Effect (95% CI)",
title = "Conditional Treatment Effect Across Baseline Depression"
) +
theme_minimal()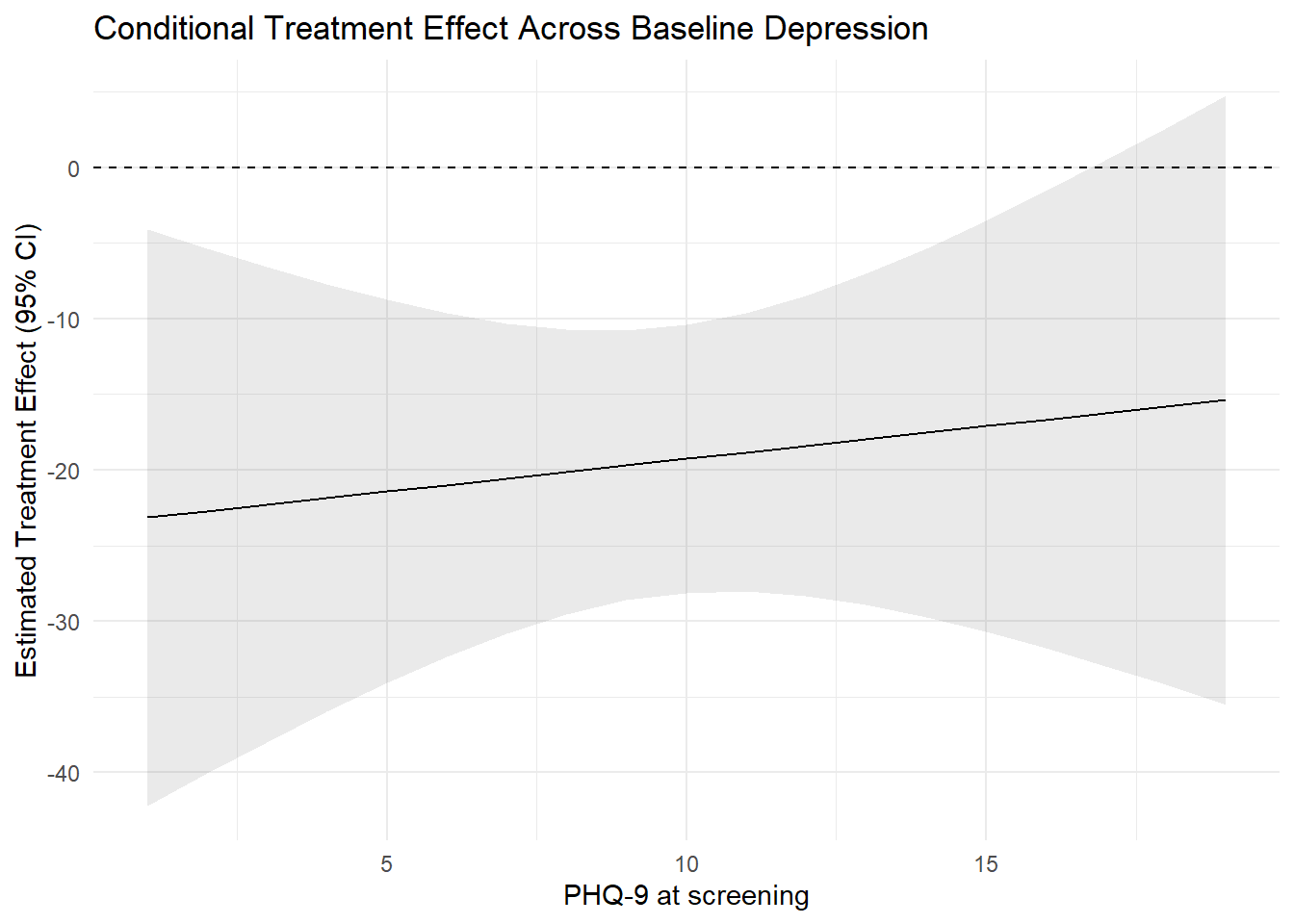
8.4 Non-linear interaction (Spline)
So far it does not seem like there is a substantial effect of baseline PHQ-9 scores on the treatment effect. However, it is possible that the effect varies non-linearly. We can investigate this by modeling PHQ-9 scores using a natural cubic spline.
library(splines)
library(patchwork)
# Fit spline regression
mod_lm_int_spline <- lm(
lsas_post ~ trt * ns(phq9_screen, df = 4),
data = df_red
)
# Plot predicted outcomes
p_spline_trend <- plot_predictions(
mod_lm_int_spline,
by = c("phq9_screen", "trt")
) +
geom_rug(
data = df_red,
aes(x = phq9_screen, y = lsas_post),
alpha = 0.25, position = "jitter", sides = "b"
) +
geom_point(
data = df_red,
aes(phq9_screen, lsas_post, group = trt, color = trt),
alpha = 0.5
) +
labs(
x = "PHQ-9 at screening",
y = "Estimated Mean LSAS (95% CI)",
color = "Treatment group",
fill = "Treatment group",
title = "Predicted Outcomes"
) +
theme_minimal()
# Plot treatment effects
p_spline_es <- plot_comparisons(
mod_lm_int_spline,
variables = "trt",
by = "phq9_screen"
) +
geom_hline(yintercept = 0, linetype = "dashed") +
labs(
x = "PHQ-9 at screening",
y = "Estimated Treatment Effect (95% CI)",
color = "Treatment group",
fill = "Treatment group",
title = "Treatment Effect"
) +
theme_minimal()
# Combine plots
(p_spline_trend + p_spline_es & theme(legend.position = "bottom")) +
plot_layout(guides = "collect") +
plot_annotation(
title = paste(
"Spline Interaction Model: Treatment Effect",
"Varies Non-linearly with PHQ-9"
)
)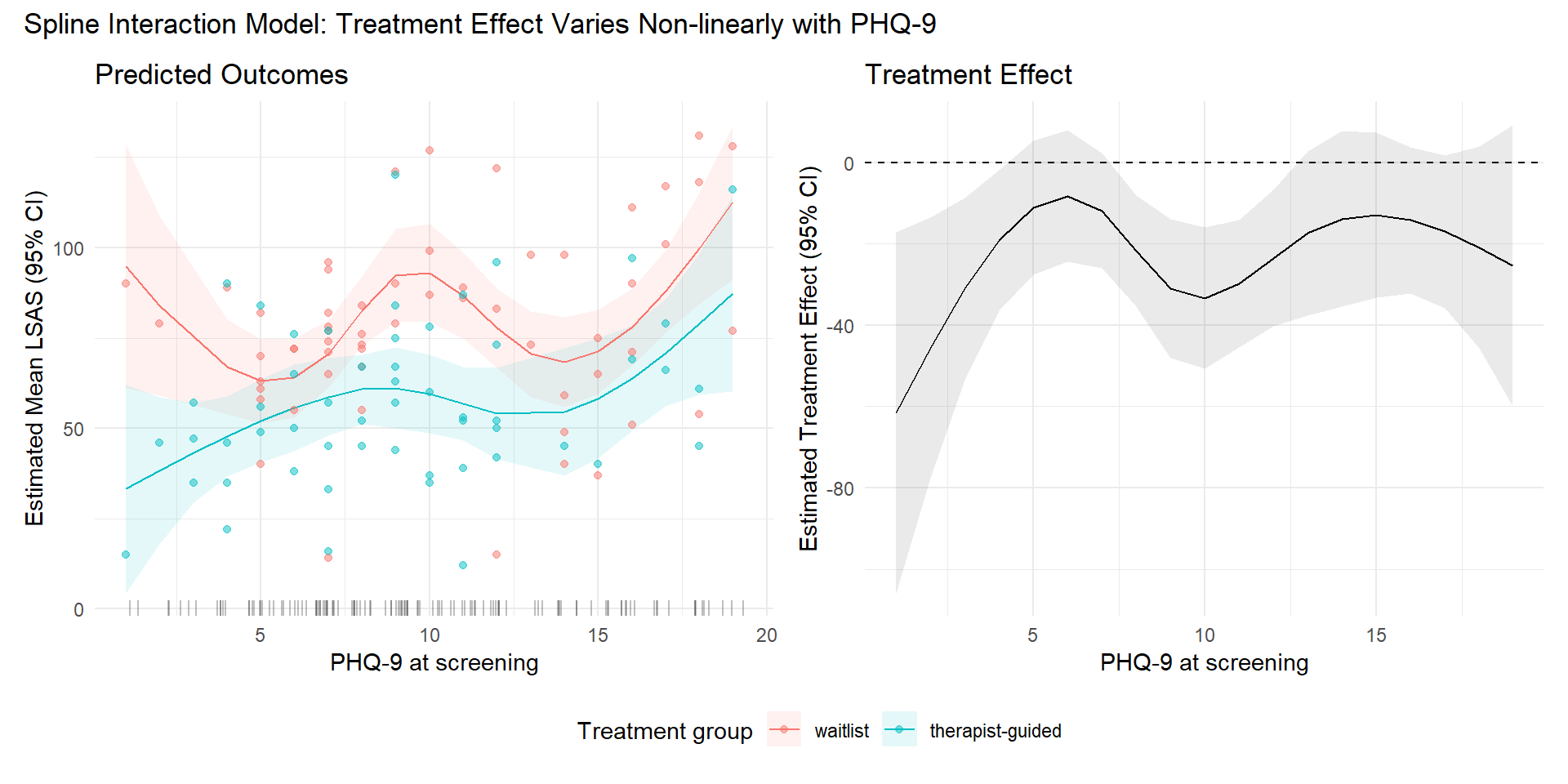
Do these curves make sense? Do we believe that the relationships are this wiggly? We should consider potential overfitting and fit a more constrained spline (e.g., changing df to 2 or 3)
8.5 Slightly More Complicated Hypothesis & Standardized Effects
We can estimate many other useful contrasts from the fitted model. For example, we might compare the treatment effect at PHQ-9 = 1 versus PHQ-9 = 15. The call below computes the difference between the two treatment effects at those PHQ-9 levels and then standardizes it by the baseline LSAS SD.
sd_baseline <- sd(df_red$lsas_screen, na.rm = TRUE)
avg_comparisons(
mod_lm_int_spline,
variables = "trt",
by = "phq9_screen",
newdata = datagrid(phq9_screen = c(1, 15)),
hypothesis = ~ reference | contrast,
transform = \(x) x / sd_baseline,
)| Hypothesis | Estimate | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|
| Type: response | ||||
| (15) - (1) | 2.7 | 0.06 | -0.113 | 5.51 |
Estimated standardized difference in treatment effects between PHQ-9 = 1 and 15 is 2.7 SD units (95% CI: [-0.133, 5.51]).
Note: The order of values in newdata determines the direction of the contrast when using hypothesis = ~ reference | contrast (the result is contrast minus reference). Swap the order if you prefer the opposite difference.
We could also compare the treatment effect at each level of PHQ-9 screen to the mean of all effects
cmp_meandev <- avg_comparisons(
mod_lm_int_spline,
variables = "trt",
by = "phq9_screen",
transform = \(x) x / sd_baseline,
hypothesis = ~ meandev | contrast
)
cmp_meandev| Hypothesis | Estimate | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|
| Type: response | ||||
| (1) - Mean | -2.1030 | 0.0642 | -4.3298 | 0.124 |
| (2) - Mean | -1.2218 | 0.1177 | -2.7522 | 0.309 |
| (3) - Mean | -0.4127 | 0.4150 | -1.4052 | 0.580 |
| (4) - Mean | 0.2518 | 0.5257 | -0.5258 | 1.029 |
| (5) - Mean | 0.6995 | 0.1063 | -0.1493 | 1.548 |
| (6) - Mean | 0.8582 | 0.0662 | -0.0574 | 1.774 |
| (7) - Mean | 0.6557 | 0.1278 | -0.1883 | 1.500 |
| (8) - Mean | 0.1087 | 0.7914 | -0.6970 | 0.914 |
| (9) - Mean | -0.4089 | 0.4013 | -1.3636 | 0.546 |
| (10) - Mean | -0.5393 | 0.2559 | -1.4698 | 0.391 |
| (11) - Mean | -0.3450 | 0.3905 | -1.1324 | 0.442 |
| (12) - Mean | 0.0068 | 0.9870 | -0.8116 | 0.825 |
| (13) - Mean | 0.3487 | 0.4958 | -0.6546 | 1.352 |
| (14) - Mean | 0.5461 | 0.3170 | -0.5234 | 1.616 |
| (15) - Mean | 0.5958 | 0.2276 | -0.3721 | 1.564 |
| (16) - Mean | 0.5273 | 0.2014 | -0.2816 | 1.336 |
| (17) - Mean | 0.3701 | 0.3926 | -0.4783 | 1.219 |
| (18) - Mean | 0.1539 | 0.8073 | -1.0826 | 1.390 |
| (19) - Mean | -0.0919 | 0.9212 | -1.9131 | 1.729 |
Let’s plot it and compare it to our previous treatment effect vs. PHQ-9 figure.
p_spline_meandev <- cmp_meandev |>
mutate(phq9_screen = row_number() + 1) |>
ggplot(aes(x = phq9_screen, y = estimate)) +
geom_line() +
geom_hline(yintercept = 0, linetype = "dashed") +
geom_ribbon(aes(ymin = conf.low, ymax = conf.high), alpha = 0.15) +
labs(
title = "Deviations from Mean Effect",
x = "PHQ-9 at screening",
y = "Deviation from mean effect (SD units, 95% CI)"
) +
theme_minimal()
p_spline_es + p_spline_meandev +
plot_annotation(
title = paste(
"Spline Interaction Model: Treatment Effect",
"Varies Non-linearly with PHQ-9"
)
)Ignoring unknown labels:
• colour : "Treatment group"
• fill : "Treatment group"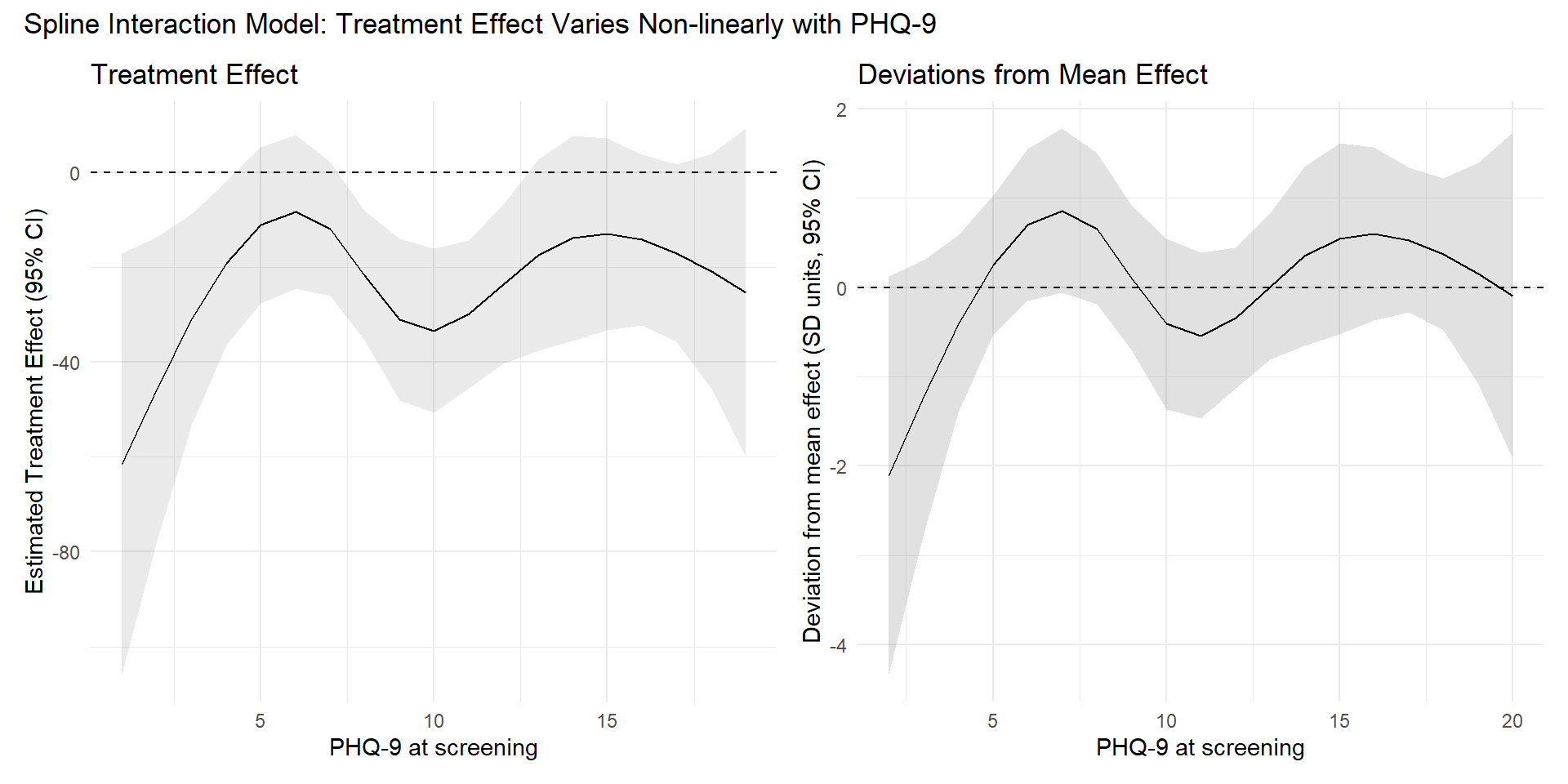
The figure above illustrates point that the difference between “significant” and “not significant” is not itself statistically significant. In the left panel, the 95% CI for the treatment effect crosses 0 at some PHQ-9 values and not at others; this alone does not imply that the formal contrast between two treatment effects (e.g., PHQ-9 = 1 vs 15) is significant. The right panel shows that the direct comparison can be non-significant even when individual CIs differ in whether they include 0.
9 Summary
In this chapter you learned:
- how to fit regression models with one binary and one numeric predictor
- how conditional effects are interpreted when a moderator is continuous
- the difference between additive (parallel slopes) and interaction models
- how interactions change coefficient interpretation
- how to estimate conditional and marginal effects using
marginaleffects - how to visualize interactions using predicted values and treatment-effect curves
This chapter continues the foundation for working with multiple predictors, including both categorical and numeric variables.