library(tidyverse)
library(broom)
library(quantreg)
library(marginaleffects)
library(here)Chapter 3: Regression with a Categorical Predictor
1 Introduction
In the previous chapter, we studied regression with a binary predictor.
In this chapter, we extend that framework to a categorical predictor with three levels, called trt.
This is a common situation in experiments with more than two arms (e.g., waitlist vs two treatment variants). The main difference compared to the binary case is that the model now estimates:
- the expected outcome in a reference group
- the difference between each other group and the reference group
You will learn:
- how categorical predictors are represented using factors in R
- how to interpret intercepts and coefficients with 3+ groups
- how to obtain predicted values for each group
- how to compute and visualize pairwise group differences
- how linear (mean) and quantile (median) regression handle categorical predictors
2 Packages and Data
2.1 Load data
df_clean <- read_csv2(here("data", "steps_clean.csv"))ℹ Using "','" as decimal and "'.'" as grouping mark. Use `read_delim()` for more control.Rows: 181 Columns: 37
── Column specification ────────────────────────────────────────────────────────
Delimiter: ";"
chr (1): trt
dbl (34): id, lsas_screen, gad_screen, phq9_screen, bbq_screen, scs_screen, ...
num (2): dmrsodf_screen, dmrsodf_post
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.We will again use the treatment indicator variable trt as our predictor. In the original data, this indicator has three levels:
table(df_clean$trt)
self-guided therapist-guided waitlist
61 60 60 To make sure R treats trt as a categorical predictor, we convert it to a factor and choose a reference level.
As in the last chapter, we set waitlist as the reference group.
df_clean <- df_clean |>
mutate(trt = factor(trt,
levels = c("waitlist", "therapist-guided", "self-guided")
))
levels(df_clean$trt)[1] "waitlist" "therapist-guided" "self-guided" Now we have the dummy coding:
contrasts(df_clean$trt) therapist-guided self-guided
waitlist 0 0
therapist-guided 1 0
self-guided 0 13 Regression with a Categorical Predictor
With a categorical predictor with three levels, the regression model estimates:
- the expected outcome in the reference group
- the difference between each of the other groups and the reference group
3.1 Model formula
lsas_post ~ trtThe R syntax is the same as with the binary predictor. However, when trt has three levels, R uses dummy coding by default (also called treatment coding).
If waitlist is the reference group, the linear regression model can be written:
\[ \mathbb{E}(LSAS_{post} \mid trt) = \beta_0 + \beta_1 \, \mathbf{1}\{trt = \text{therapist-guided}\} + \beta_2 \, \mathbf{1}\{trt = \text{self-guided}\} \]
Where:
- \(\beta_0\) is the expected outcome in the reference group (
waitlist) - \(\beta_1\) is the mean difference between therapist-guided and waitlist
- \(\beta_2\) is the mean difference between self-guided and waitlist
- \(\mathbf{1}\{\cdot\}\) is the indicator function which is 1 if the condition holds and 0 otherwise
| trt | Predicted value |
|---|---|
| waitlist | \(\beta_0\) |
| therapist-guided | \(\beta_0 + \beta_1\) |
| self-guided | \(\beta_0 + \beta_2\) |
So the coefficients compare each treatment arm to the reference group.
4 Linear Regression (Mean Differences)
4.1 Fit the model
mod_lm <- lm(lsas_post ~ trt, data = df_clean)
summary(mod_lm)
Call:
lm(formula = lsas_post ~ trt, data = df_clean)
Residuals:
Min 1Q Median 3Q Max
-64.448 -14.912 -1.352 14.088 62.648
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 78.448 3.062 25.623 < 2e-16 ***
trttherapist-guided -21.096 4.409 -4.785 3.76e-06 ***
trtself-guided -13.536 4.349 -3.113 0.00218 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 23.32 on 166 degrees of freedom
(12 observations deleted due to missingness)
Multiple R-squared: 0.1248, Adjusted R-squared: 0.1143
F-statistic: 11.84 on 2 and 166 DF, p-value: 1.561e-05(Intercept): wait-list meantrttherapist-guided: mean difference for the therapist-guided vs wait-listtrtself-guided: mean difference for the self-guided group vs wait-list
We can verify group means directly:
df_clean |>
group_by(trt) |>
summarize(mean_lsas = mean(lsas_post, na.rm = TRUE))| trt | mean_lsas |
|---|---|
| waitlist | 78.44828 |
| therapist-guided | 57.35185 |
| self-guided | 64.91228 |
4.2 Relationship to one-way ANOVA
A linear regression with one categorical predictor (3+ groups) is equivalent to a one-way ANOVA:
anova(mod_lm)| Df | Sum Sq | Mean Sq | F value | Pr(>F) | |
|---|---|---|---|---|---|
| trt | 2 | 12873.37 | 6436.685 | 11.83959 | 1.56e-05 |
| Residuals | 166 | 90247.22 | 543.658 | NA | NA |
The ANOVA table tests whether there is evidence that at least one group mean differs.
5 Quantile Regression (Quantile Differences)
5.1 Model formulation
For quantile regression at quantile \(\tau\):
\[ Q_{LSAS_{post}}(\tau) = \beta_0(\tau) + \beta_1(\tau) \, \mathbf{1}\{trt = \text{therapist-guided}\} + \beta_2(\tau) \, \mathbf{1}\{trt = \text{self-guided}\} \]
- \(\beta_0(\tau)\) = \(\tau\)-th quantile in waitlist
- \(\beta_1(\tau)\) = \(\tau\)-th quantile difference (therapist-guided − waitlist)
- \(\beta_2(\tau)\) = \(\tau\)-th quantile difference (self-guided − waitlist)
5.2 Fit a median regression
mod_rq <- rq(lsas_post ~ trt, tau = 0.5, data = df_clean)
summary(mod_rq)
Call: rq(formula = lsas_post ~ trt, tau = 0.5, data = df_clean)
tau: [1] 0.5
Coefficients:
coefficients lower bd upper bd
(Intercept) 77.00000 71.70143 84.59713
trttherapist-guided -24.00000 -31.74701 -16.62650
trtself-guided -13.00000 -18.30306 -2.13129Again, the interpretation of the coefficients is the same as in linear regression, but now refers to quantiles.
Verify group medians directly:
df_clean |>
group_by(trt) |>
summarize(median_lsas = median(lsas_post, na.rm = TRUE))| trt | median_lsas |
|---|---|
| waitlist | 77.0 |
| therapist-guided | 52.5 |
| self-guided | 64.0 |
6 Predictions Using marginaleffects
As in the binary predictor case, marginaleffects makes it easy to get:
- predicted values for each group
- uncertainty (SEs and CIs)
- group differences as contrasts
6.1 Predicted means by group (linear regression)
avg_predictions(mod_lm, by = "trt")| trt | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| waitlist | 78.4 | 3.06 | 25.6 | <0.001 | 72.4 | 84.4 |
| therapist-guided | 57.4 | 3.17 | 18.1 | <0.001 | 51.1 | 63.6 |
| self-guided | 64.9 | 3.09 | 21.0 | <0.001 | 58.9 | 71.0 |
6.2 Predicted medians by group (quantile regression)
avg_predictions(mod_rq, by = "trt")| trt | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| waitlist | 77 | 3.36 | 22.9 | <0.001 | 70.4 | 83.6 |
| therapist-guided | 53 | 4.07 | 13.0 | <0.001 | 45.0 | 61.0 |
| self-guided | 64 | 3.96 | 16.2 | <0.001 | 56.2 | 71.8 |
6.3 Group differences as contrasts
6.3.1 Mean differences (linear regression)
With three groups, there is no single “treatment − control” difference. Instead, we usually calculate the pairwise mean differences between all groups.
cmp <- avg_comparisons(mod_lm, variables = list("trt" = "pairwise"))
cmp| Contrast | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| self-guided - therapist-guided | 7.56 | 4.43 | 1.71 | 0.08773 | -1.12 | 16.24 |
| self-guided - waitlist | -13.54 | 4.35 | -3.11 | 0.00185 | -22.06 | -5.01 |
| therapist-guided - waitlist | -21.10 | 4.41 | -4.78 | < 0.001 | -29.74 | -12.45 |
6.3.2 Median differences (quantile regression)
cmp_med <- avg_comparisons(mod_rq, variables = list("trt" = "pairwise"))
cmp_med| Contrast | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| self-guided - therapist-guided | 11 | 5.67 | 1.94 | 0.0525 | -0.12 | 22.12 |
| self-guided - waitlist | -13 | 5.19 | -2.50 | 0.0123 | -23.18 | -2.82 |
| therapist-guided - waitlist | -24 | 5.28 | -4.55 | <0.001 | -34.34 | -13.66 |
7 Visualization
7.1 Group means with confidence intervals
plot_predictions(mod_lm, by = "trt") +
labs(
x = "Treatment group",
y = "Estimated Mean LSAS (95% CI)",
title = "Estimated Group Means from Linear Regression"
) +
theme_minimal()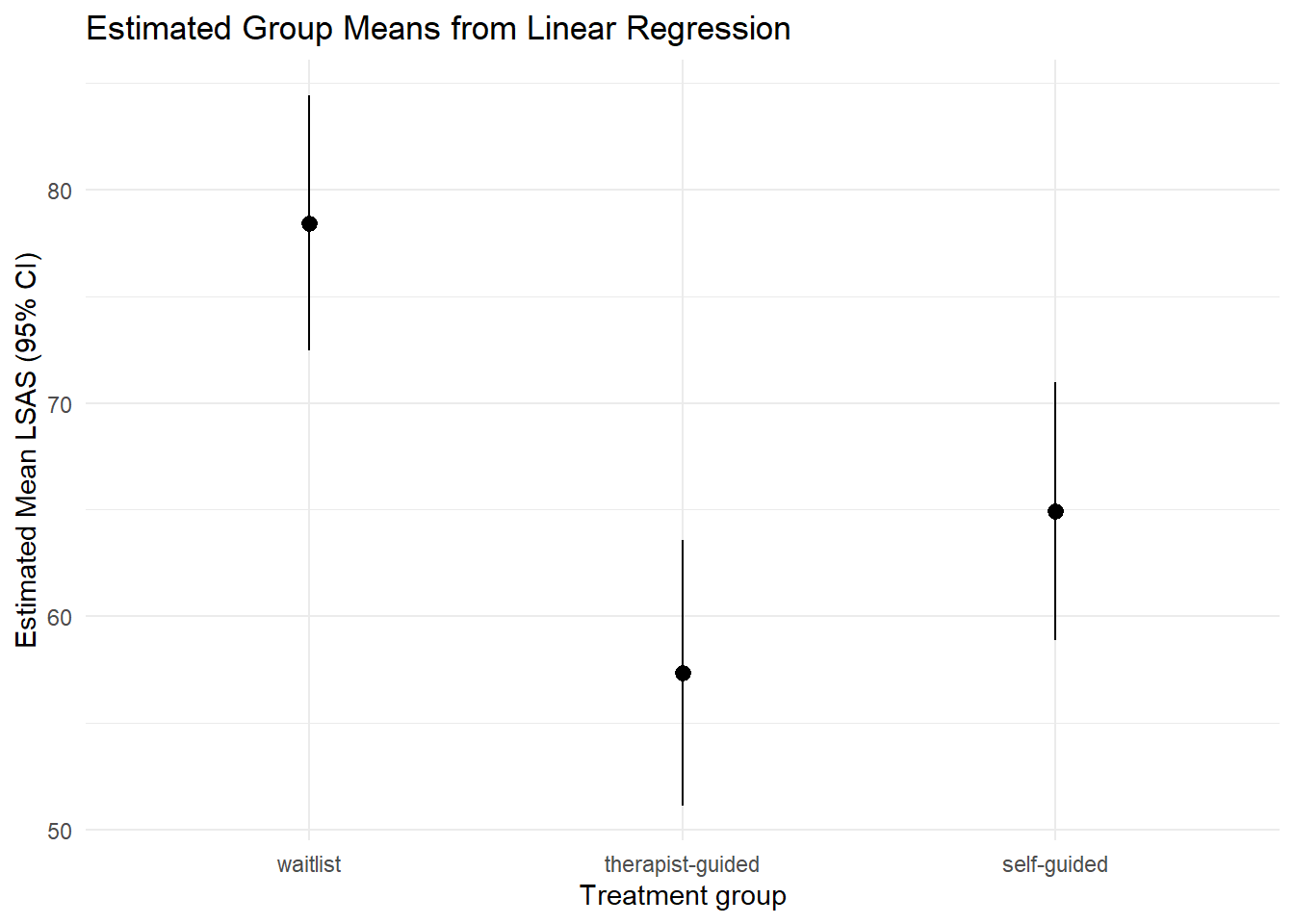
7.2 Group medians with confidence intervals
plot_predictions(mod_rq, by = "trt") +
labs(
x = "Treatment group",
y = "Estimated Median LSAS (95% CI)",
title = "Estimated Group Medians from Quantile Regression"
) +
theme_minimal()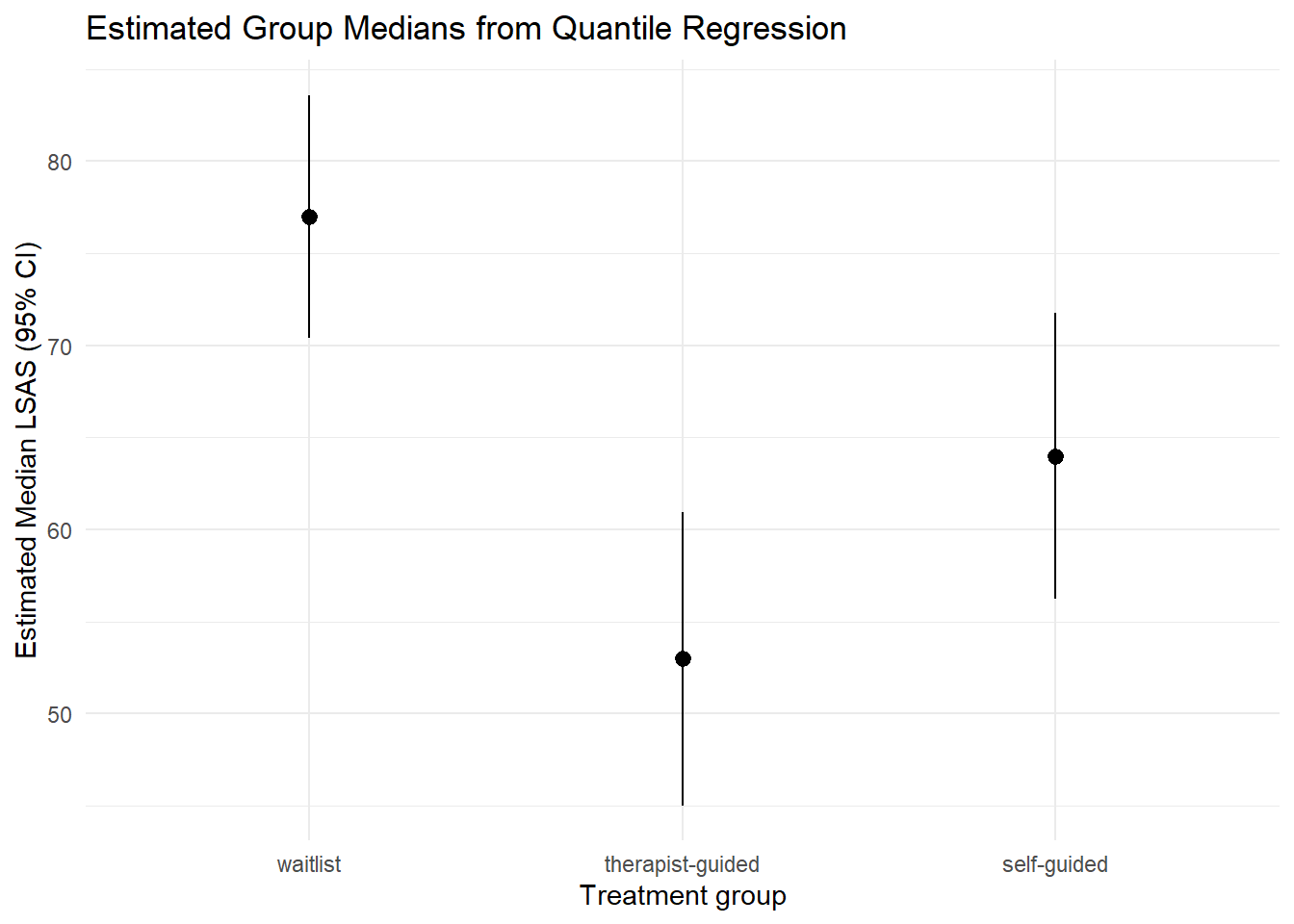
7.3 Mean differences (pairwise) with confidence intervals
cmp |>
ggplot(aes(y = contrast, x = estimate)) +
geom_vline(xintercept = 0, linetype = "dashed") +
geom_point(size = 3) +
geom_errorbar(aes(xmin = conf.low, xmax = conf.high), width = 0.1) +
labs(
y = "",
x = "Estimated Mean Difference (95% CI)",
title = "Pairwise Mean Differences (Linear Regression)"
) +
theme_minimal()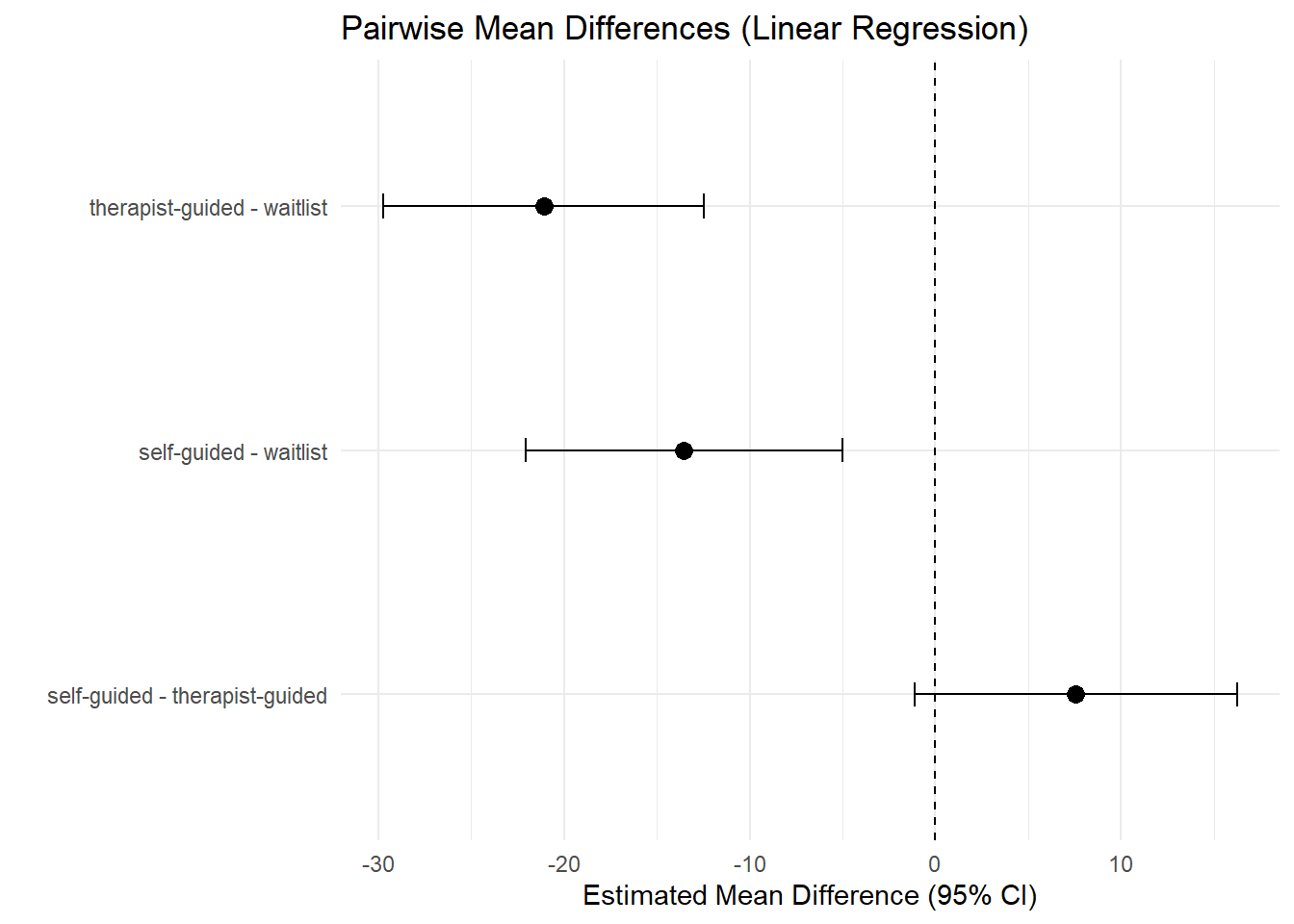
7.4 Median differences (pairwise) with confidence intervals
ggplot(cmp_med, aes(y = contrast, x = estimate)) +
geom_vline(xintercept = 0, linetype = "dashed") +
geom_point(size = 3) +
geom_errorbar(aes(xmin = conf.low, xmax = conf.high), width = 0.1) +
labs(
y = "",
x = "Estimated Median Difference (95% CI)",
title = "Pairwise Median Differences (Quantile Regression)"
) +
theme_minimal()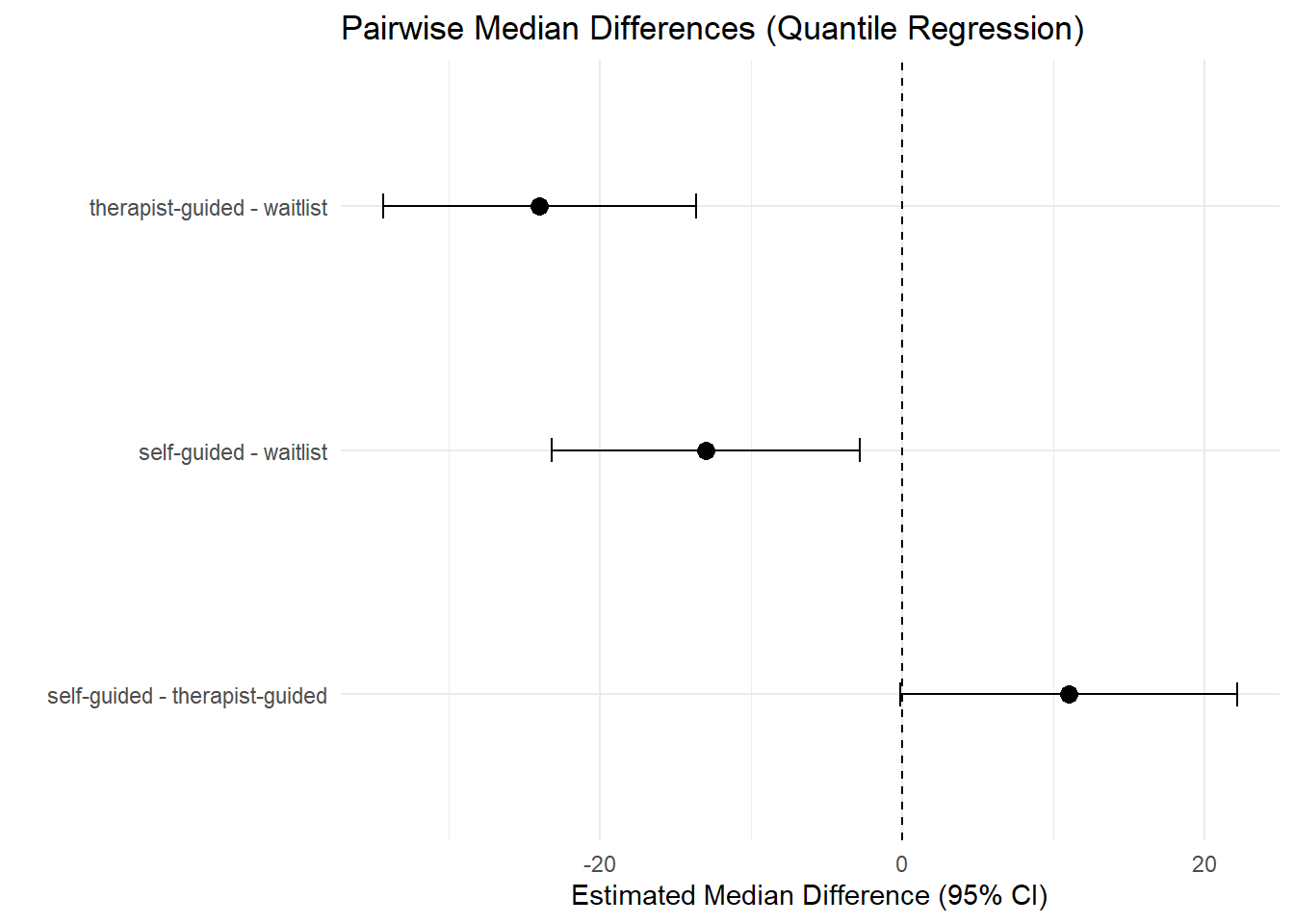
7.5 Ratio of Means and Other Hypotheses
As before, we can answer a range of other hypotheses. For example, the ratio of group outcomes.
avg_comparisons(
mod_lm,
variables = list("trt" = "pairwise"),
comparison = "ratio"
)| Contrast | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| mean(self-guided) / mean(therapist-guided) | 1.132 | 0.0826 | 13.7 | <0.001 | 0.970 | 1.294 |
| mean(self-guided) / mean(waitlist) | 0.827 | 0.0509 | 16.3 | <0.001 | 0.728 | 0.927 |
| mean(therapist-guided) / mean(waitlist) | 0.731 | 0.0495 | 14.8 | <0.001 | 0.634 | 0.828 |
We could also compare each treatment group to the grand mean, instead of contrasting them with a reference group (the waitlist).
avg_predictions(
mod_lm,
by = "trt",
hypothesis = ~meandev
)| Hypothesis | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| (waitlist) - Mean | 11.54 | 2.52 | 4.583 | <0.001 | 6.61 | 16.48 |
| (therapist-guided) - Mean | -9.55 | 2.56 | -3.725 | <0.001 | -14.58 | -4.53 |
| (self-guided) - Mean | -1.99 | 2.53 | -0.787 | 0.431 | -6.95 | 2.97 |
8 Summary
In this chapter you learned:
- how to fit regression models with a categorical predictor (3 levels)
- how dummy coding works (reference group + differences)
- how linear regression estimates mean differences across groups
- how quantile regression estimates quantile differences across groups
- how to extract predicted values and contrasts using
marginaleffects - how to visualize group means/medians and pairwise differences with confidence intervals