library(tidyverse)
library(broom)
library(quantreg)
library(marginaleffects)
library(here)Chapter 2: Regression with a Binary Predictor
1 Introduction
In the previous chapter, we studied intercept-only regression models, which estimate a single summary of the outcome (a mean or a quantile).
In this chapter, we extend that framework by adding one binary predictor, called trt.
This is the simplest possible comparative regression model and corresponds to:
- a two-sample t-test in linear regression
- a comparison of quantiles in quantile regression
This chapter introduces how regression models encode group differences.
You will learn:
- how binary predictors enter regression models
- how to interpret intercepts and slopes with binary variables
- how group means and differences are estimated
- how to obtain predicted values for each group
- how to visualize group comparisons
- how linear and quantile regression differ in interpretation
2 Packages and Data
We use the same packages as before.
2.1 Load data
We load the same data as before.
df_clean <- read_csv2(here("data", "steps_clean.csv"))We will use the treatment indicator variable trt as our predictor. In the original data, this indicator has three levels:
levels(as.factor(df_clean$trt))[1] "self-guided" "therapist-guided" "waitlist" To get a binary variable (i.e., one with only two levels), we subset the data to include only participants in the therapist-guided or wait-list conditions. We’ll call this reduced dataset df_red.
df_red <- filter(df_clean, trt %in% c("therapist-guided", "waitlist"))We transform the trt variable to a factor, and make sure that the wait-list condition is used as the reference.
df_red <- df_red |>
mutate(trt = factor(trt, levels = c("waitlist", "therapist-guided")))By listing wait-list as the first level, it becomes the reference level. We can check how R converts the factor to regression contrasts using contrasts.
contrasts(df_red$trt) therapist-guided
waitlist 0
therapist-guided 13 Regression with a Binary Predictor
With a binary predictor, the regression model estimates two quantities:
- the expected outcome in the reference group
- the difference between groups
3.1 Model formula
Here is the R syntax for the regression model for the LSAS post-treatment score, lsas_post, including only an intercept and the trt variable as a predictor (the intercept ~1 from the last model is implied, but can also be written out).
lsas_post ~ trtlsas_post ~ 1 + trtThis corresponds to the linear regression model for the conditional mean:
\[ \mathbb{E}(LSAS_{post} \mid trt) = \beta_0 + \beta_1\, trt \]
Where:
- \(\beta_0\) is the expected outcome when
trt = 0(waitlist group, since this is coded as the reference level) - \(\beta_1\) is the mean difference between the therapist-guided and waitlist groups (therapist-guided minus waitlist)
For individual observations:
\[ LSAS_{post,i} = \beta_0 + \beta_1\, trt_i + \epsilon_i \]
3.2 Interpretation of coefficients
| trt | Predicted value |
|---|---|
| 0 (waitlist) | \(\beta_0\) |
| 1 (therapist-guided) | \(\beta_0 + \beta_1\) |
Thus:
- \(\beta_0\) = mean (or quantile) in the control group
- \(\beta_1\) = difference in mean (or quantile) between the groups (i.e., treatment effect)
4 Linear Regression (Mean Differences)
4.1 Fit the model
mod_lm <- lm(lsas_post ~ trt, data = df_red)
summary(mod_lm)
Call:
lm(formula = lsas_post ~ trt, data = df_red)
Residuals:
Min 1Q Median 3Q Max
-64.448 -13.448 -1.948 12.624 62.648
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 78.448 3.201 24.508 < 2e-16 ***
trttherapist-guided -21.096 4.610 -4.576 1.25e-05 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 24.38 on 110 degrees of freedom
(8 observations deleted due to missingness)
Multiple R-squared: 0.1599, Adjusted R-squared: 0.1523
F-statistic: 20.94 on 1 and 110 DF, p-value: 1.25e-05- Intercept: estimated mean LSAS score in the control group
- trt coefficient: estimated mean difference (therapist-guided minus waitlist). A negative value indicates lower mean LSAS at post in the therapist-guided group compared to the waitlist group. We can see that the mean value of LSAS at post was 21 point lower in the therapist-guided group than in the waitlist group.
We can verify these directly from the data:
df_red |>
group_by(trt) |>
summarize(mean_lsas = mean(lsas_post, na.rm = TRUE))| trt | mean_lsas |
|---|---|
| waitlist | 78.44828 |
| therapist-guided | 57.35185 |
The difference between these two means equals the estimated coefficient for trt.
4.2 Relationship to the t-test
This regression model is mathematically equivalent to a two-sample t-test:
t.test(lsas_post ~ trt, data = df_red, var.equal = TRUE)
Two Sample t-test
data: lsas_post by trt
t = 4.5763, df = 110, p-value = 1.25e-05
alternative hypothesis: true difference in means between group waitlist and group therapist-guided is not equal to 0
95 percent confidence interval:
11.96065 30.23220
sample estimates:
mean in group waitlist mean in group therapist-guided
78.44828 57.35185 Regression generalizes the t-test by allowing:
- additional predictors
- non-normal outcomes
- different estimands (e.g., quantiles)
5 Quantile Regression (Quantile Differences)
5.1 Model formulation
For quantile regression at quantile \(\tau\):
\[ Q_{LSAS_{post} \mid trt}(\tau \mid trt_i) = \beta_{0}(\tau) + \beta_{1}(\tau)\, trt_i \]
- \(\beta_{0}(\tau)\) = \(\tau\)-th quantile in the control group
- \(\beta_{1}(\tau)\) = difference in quantiles between groups
5.2 Fit a median regression
mod_rq <- rq(lsas_post ~ trt, tau = 0.5, data = df_red)
summary(mod_rq)
Call: rq(formula = lsas_post ~ trt, tau = 0.5, data = df_red)
tau: [1] 0.5
Coefficients:
coefficients lower bd upper bd
(Intercept) 77.00000 71.68339 84.63323
trttherapist-guided -24.00000 -31.77207 -16.61396Interpretation is parallel to that for linear regression, but refers to quantiles instead of means. We can again verify group medians directly form the observed data:
df_red |>
group_by(trt) |>
summarize(median_lsas = median(lsas_post, na.rm = TRUE))| trt | median_lsas |
|---|---|
| waitlist | 77.0 |
| therapist-guided | 52.5 |
6 Predictions Using marginaleffects
The marginaleffects package allows us to compute group-specific predictions. This allows us to see confidence intervals for the mean (or quantile) for each group, as well as the CI for the mean (or quantile) difference between the groups.
6.1 Predicted means by group (linear regression)
avg_predictions(mod_lm, by = "trt")| trt | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| waitlist | 78.4 | 3.20 | 24.5 | <0.001 | 72.2 | 84.7 |
| therapist-guided | 57.4 | 3.32 | 17.3 | <0.001 | 50.8 | 63.9 |
This returns:
- predicted mean in each group
- standard errors
- confidence intervals for each predicted mean
6.2 Predicted medians by group (quantile regression)
avg_predictions(mod_rq, by = "trt")| trt | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| waitlist | 77 | 3.09 | 24.9 | <0.001 | 70.9 | 83.1 |
| therapist-guided | 53 | 3.71 | 14.3 | <0.001 | 45.7 | 60.3 |
6.3 Treatment effect as a contrast
avg_comparisons(mod_lm, variables = "trt")| Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|
| Type: response | |||||
| -21.1 | 4.61 | -4.58 | <0.001 | -30.1 | -12.1 |
For quantile regression:
avg_comparisons(mod_rq, variables = "trt")| Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|
| Type: response | |||||
| -24 | 4.83 | -4.97 | <0.001 | -33.5 | -14.5 |
6.4 Ratios
marginaleffects is highly flexible with regards to what hypothesis we can answer. For we example we can compare the ratio of mean outcomes.
# Mean ratio
hypotheses(
mod_lm,
hypothesis = "(`(Intercept)` + `trttherapist-guided`) / `(Intercept)` = 1"
)| Hypothesis | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| (`(Intercept)`+`trttherapist-guided`)/`(Intercept)`=1 | -0.269 | 0.0518 | -5.2 | <0.001 | -0.37 | -0.167 |
# Identical, but using b* syntax
hypotheses(
mod_lm,
hypothesis = "(b1 + b2) / b1 = 1"
)| Hypothesis | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| (b1+b2)/b1=1 | -0.269 | 0.0518 | -5.2 | <0.001 | -0.37 | -0.167 |
# Ratio of quantiles
hypotheses(
mod_rq,
hypothesis = "(b1 + b2) / b1 = 1"
)| Hypothesis | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| (b1+b2)/b1=1 | -0.312 | 0.0556 | -5.61 | <0.001 | -0.421 | -0.203 |
We can also change the reference level without having to refit the model (and much more).
avg_comparisons(
mod_lm,
variables = list(trt = c("therapist-guided", "waitlist"))
)| Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|
| Type: response | |||||
| 21.1 | 4.61 | 4.58 | <0.001 | 12.1 | 30.1 |
6.5 Standardized Effect Sizes
If we want to report standardized mean differences (Cohen’s d), we can simply scale the estimates using an appropriate standardizer, for example the standard deviation at baseline.
sd_baseline <- sd(df_red$lsas_screen, na.rm = TRUE)
# Scale each estimate
smd <- avg_comparisons(
mod_lm,
) |>
mutate(
estimate = estimate / sd_baseline,
std.error = std.error / sd_baseline,
conf.low = conf.low / sd_baseline,
conf.high = conf.high / sd_baseline
)
smd| Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|
| -1.17 | 0.256 | -4.58 | <0.001 | -1.68 | -0.671 |
# Equivalently, add standardization to hypothesis
avg_comparisons(
mod_lm,
variables = "trt",
hypothesis = "b1 / 17.97963 = 0" # sd_baseline is 17.97963
)| Hypothesis | Estimate | Std. Error | z | Pr(>|z|) | 2.5 % | 97.5 % |
|---|---|---|---|---|---|---|
| Type: response | ||||||
| b1/17.97963=0 | -1.17 | 0.256 | -4.58 | <0.001 | -1.68 | -0.671 |
7 Visualization
7.1 Group means with confidence intervals
Using marginaleffects::plot_predictions
plot_predictions(mod_lm, by = "trt") +
labs(
x = "Treatment group",
y = "Estimated Mean LSAS (95% CI)",
title = "Estimated Group Means from Linear Regression"
) +
theme_minimal()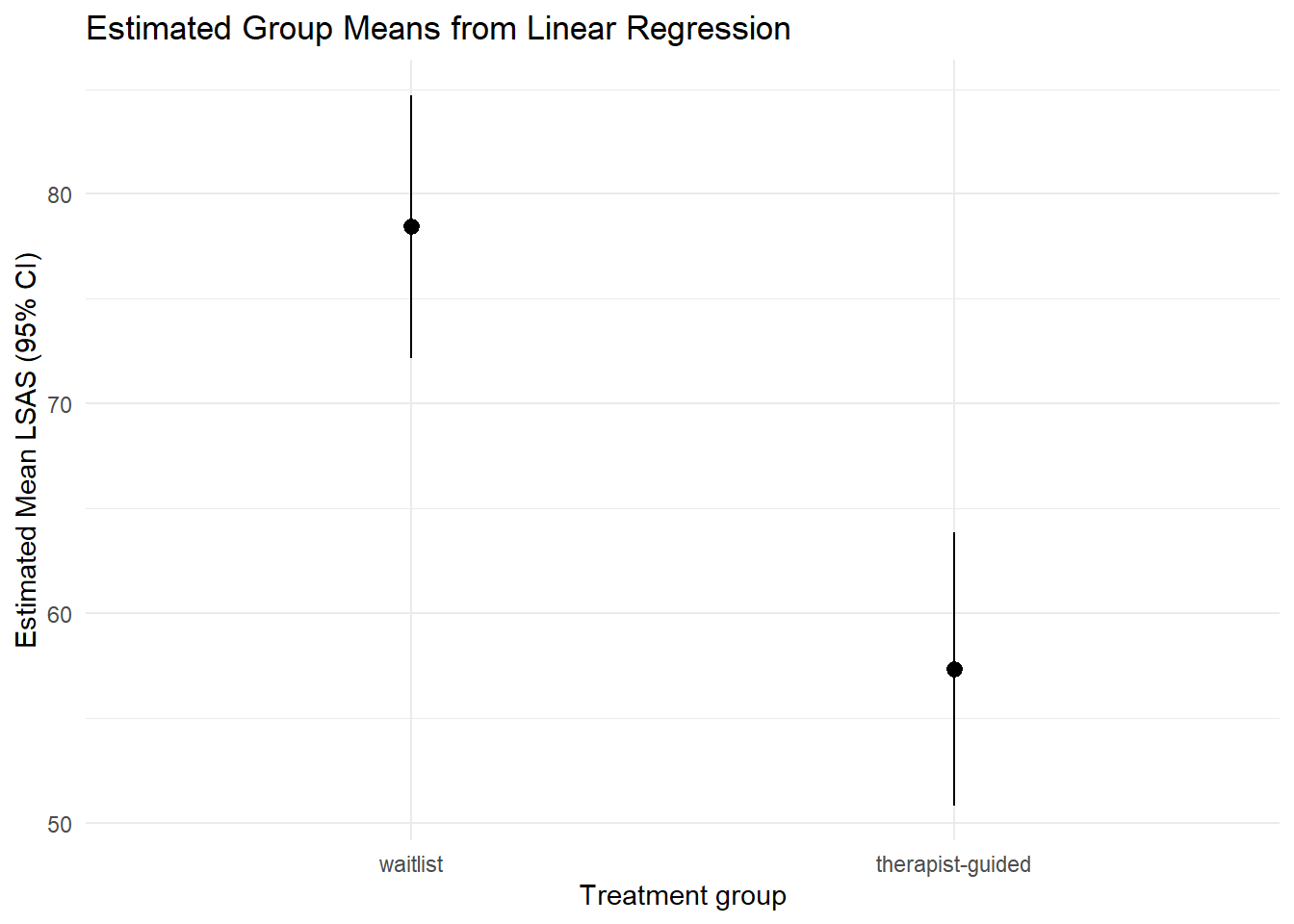
7.2 Group medians with confidence intervals
plot_predictions(mod_rq, by = "trt") +
labs(
x = "Treatment group",
y = "Estimated Median LSAS (95% CI)",
title = "Estimated Group Medians from Quantile Regression"
) +
theme_minimal()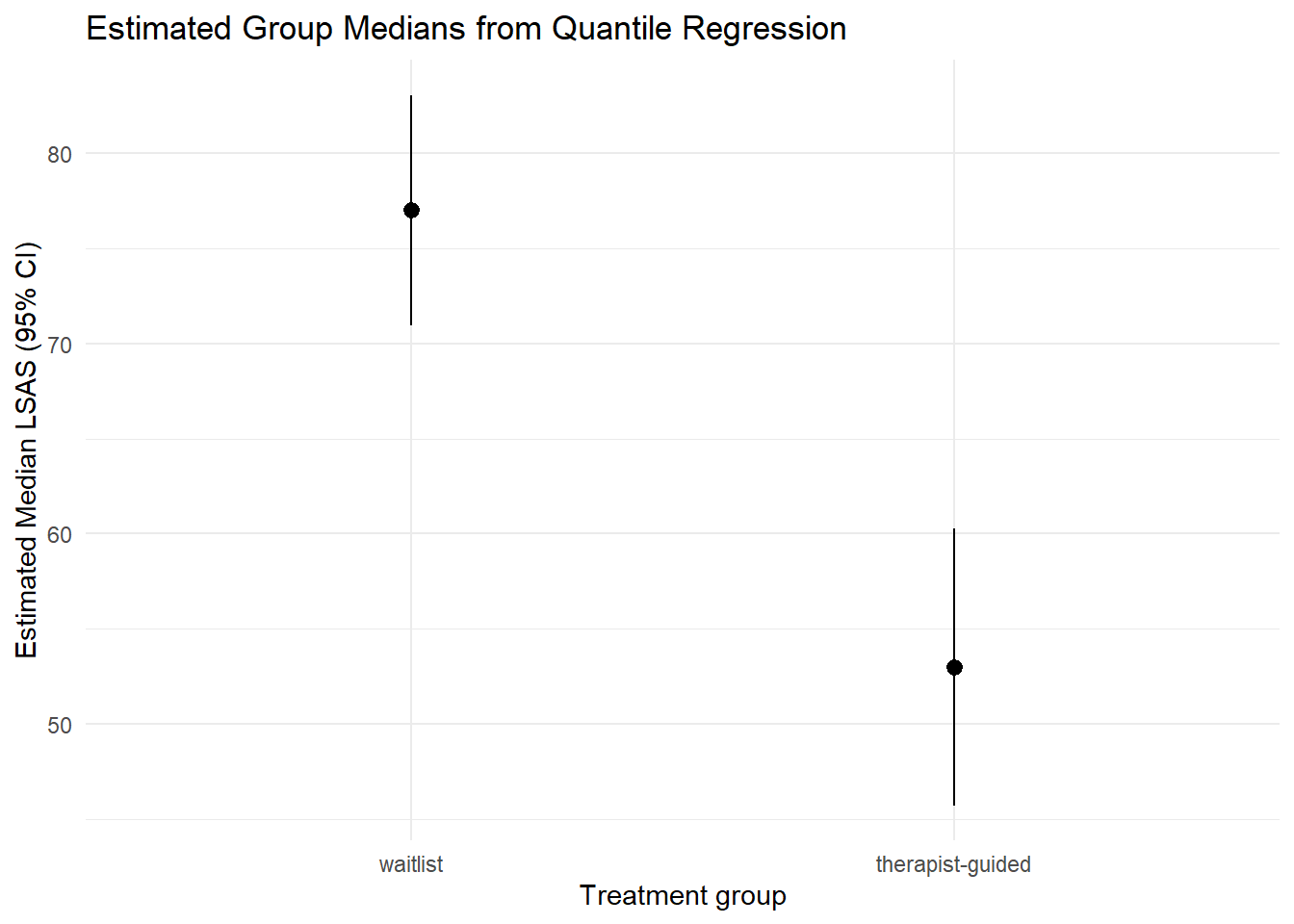
8 Summary
In this chapter you learned:
- how binary predictors enter regression models
- how intercepts and coefficients (slopes) encode group means and differences
- how linear regression estimates mean differences
- how quantile regression estimates quantile differences
- how regression relates to t-tests
- how to obtain predictions, contrasts, and confidence intervals
- how to visualize group comparisons
This chapter provides the foundation for models with categorical predictors and later numeric predictors, and multiple predictors.